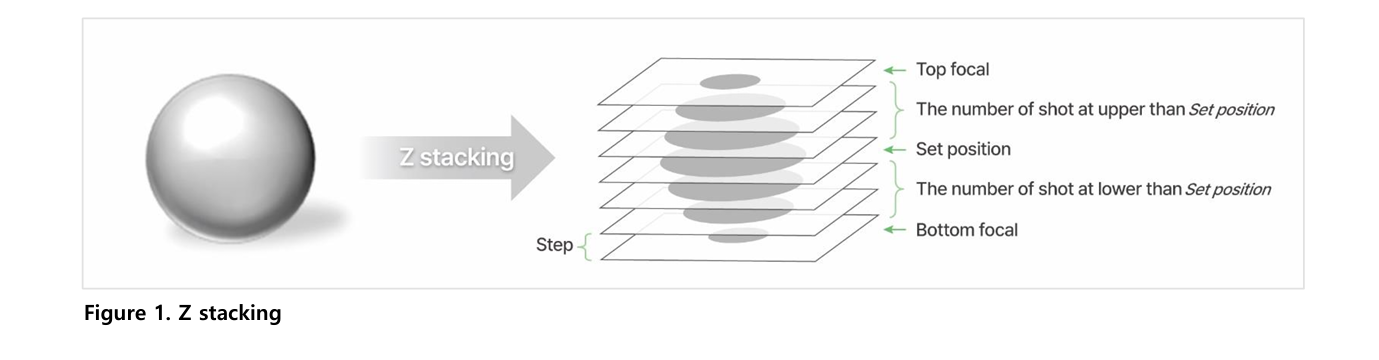

Z-stacking is a method that captures multiple images at different focal depths of a 3D sample. It is used to visualize structures that cannot be seen in a single focal plane, such as tissues within zebrafish larvae. This approach ensures detailed observation of complex three-dimensional structures.
The Celloger Mini Plus uses a motorized Z stage to capture images above and below a defined Z position. Users set the step size and number of images, and the system automatically acquires evenly spaced focal planes. This enables consistent and reproducible multi-layer imaging.
Z-projection merges multiple Z-stacked images into a single image. This reduces the need to manually analyze each focal plane and allows researchers to view 3D information at a glance. It makes interpretation faster and less labor-intensive.
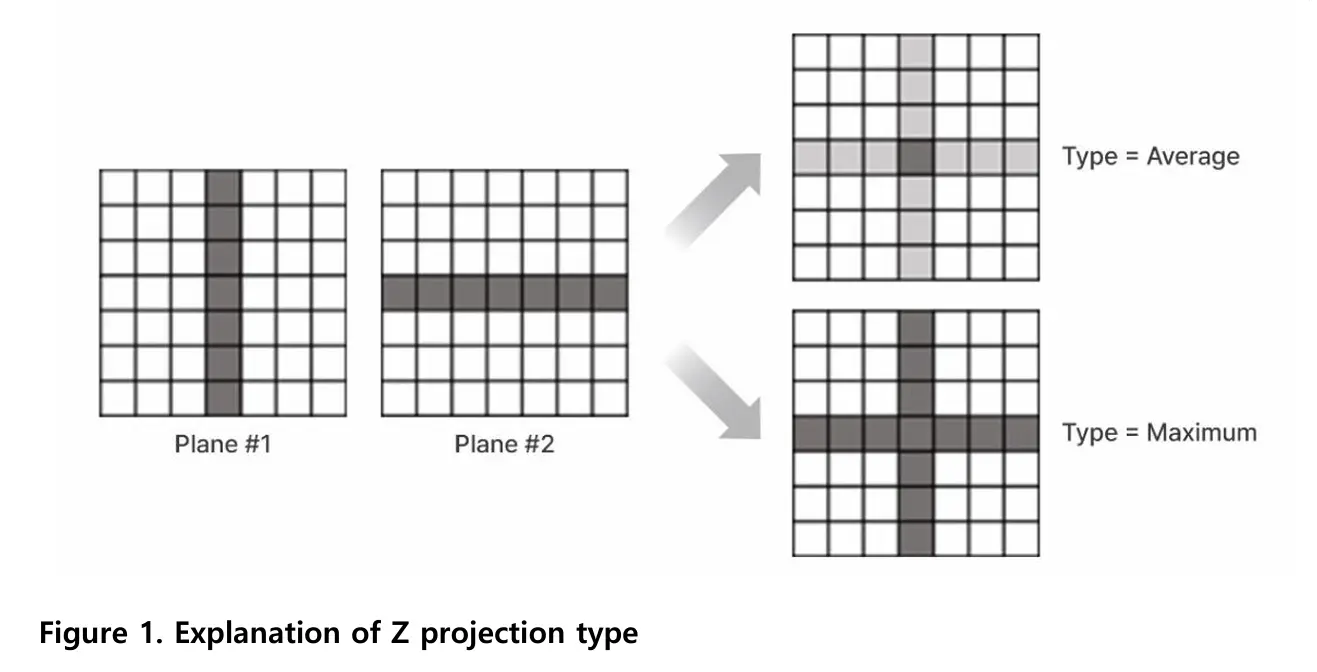
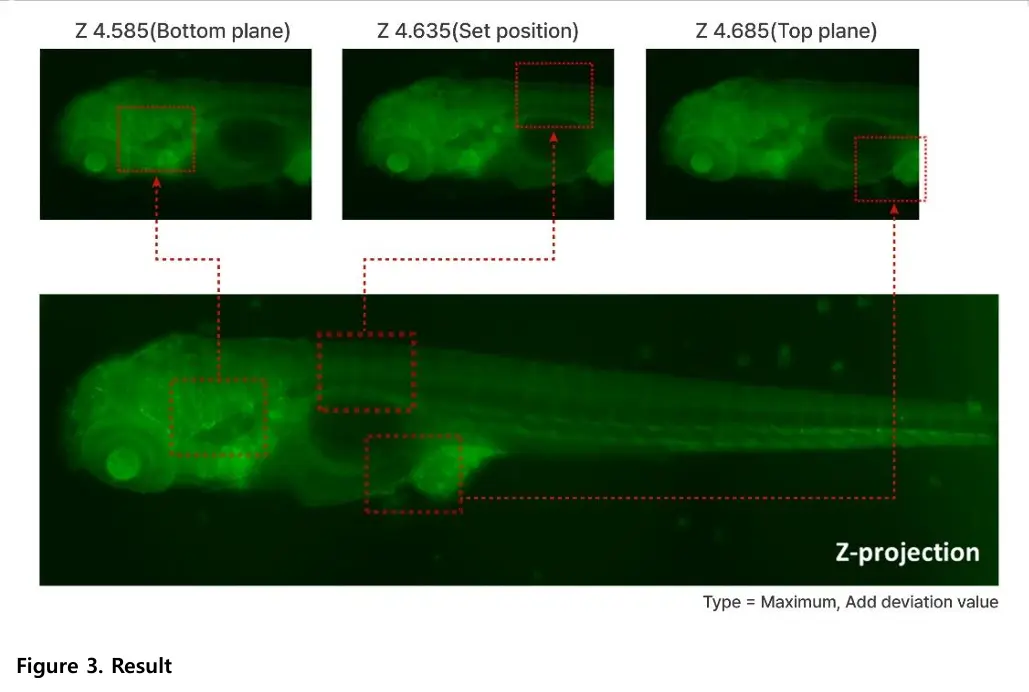
Maximum Z-projection selects the brightest pixel from all focal planes at each X-Y position. Average Z-projection calculates the mean brightness across focal planes. The optimal method depends on the sample and imaging goal.
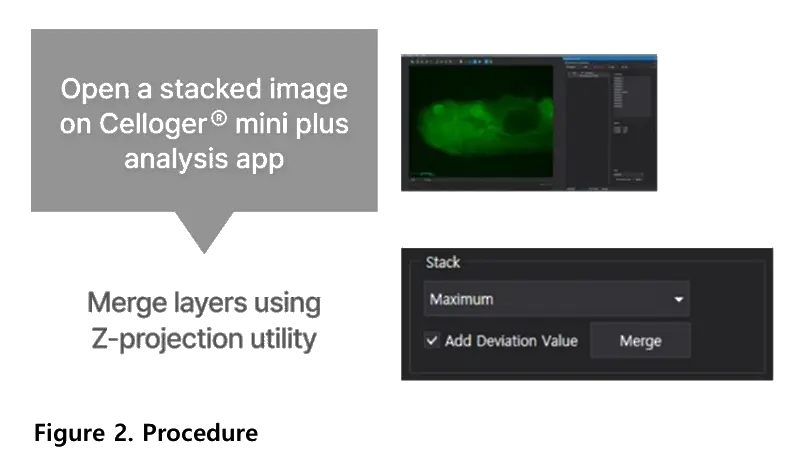
The add deviation value option enhances contrast in Z-projected images. It reinforces signal differences across focal planes, resulting in clearer visualization of fluorescent structures. This feature is available in the Celloger analysis software.
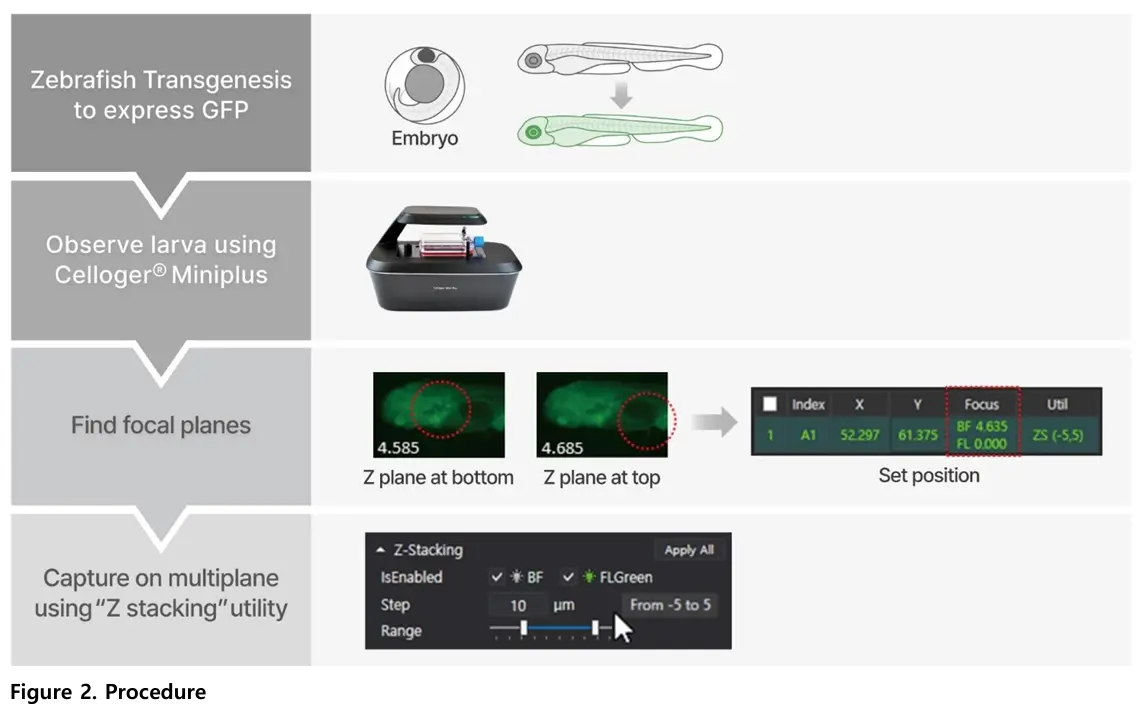
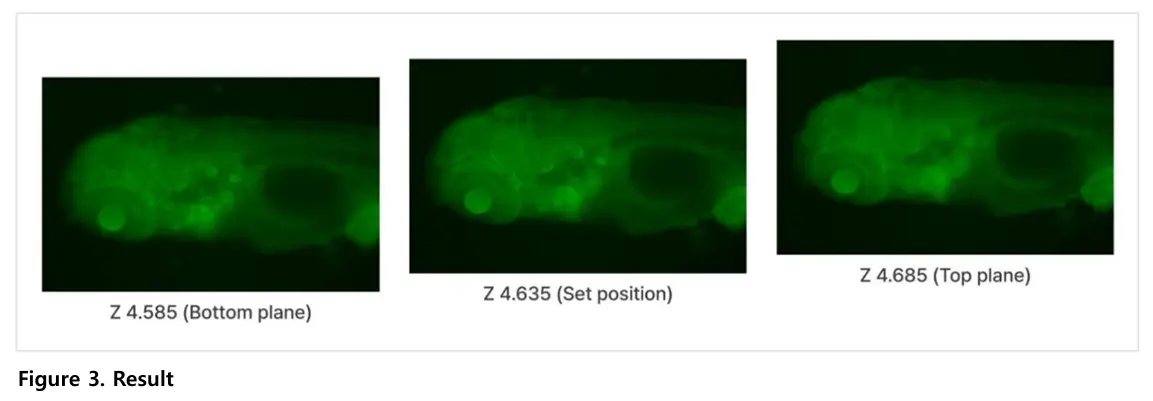
The study imaged transgenic zebrafish larvae expressing green fluorescent protein (GFP). Fluorescence imaging allowed clear visualization of internal structures across multiple focal planes.
Zebrafish larvae are three-dimensional, and different anatomical features come into focus at different depths. Capturing multiple focal planes ensures that both dorsal and ventral structures are clearly observed. This provides more complete structural information.
Yes, the application note confirms that the Celloger system can be applied to other 3D samples. These include spheroids and organoids, expanding its usability beyond zebrafish models. This demonstrates the versatility of the Celloger Mini Plus.
Celloger Mini Plus combines Z-stacking with Z-projection through its analysis software. Multi-layer fluorescence images are merged into one clear image that represents the full 3D structure. This allows researchers to assess complex morphology efficiently.
The system enables automated Z-stacking, fluorescence imaging, and easy image merging. It produces high-quality images that clearly represent three-dimensional structures. This improves data clarity while reducing manual effort during analysis.
Download the full Application Note PDF
to access the complete fluorescence Z-stacking and Z-projection workflow, including imaging settings, projection modes,
and example datasets demonstrating clear 3D visualization of GFP-expressing zebrafish larvae using the Celloger® Mini Plus.
