 Inform minimal residual disease
Inform minimal residual disease
 Post treatment monitoring
Post treatment monitoring
 Identify relapse earlier than standard of care
Identify relapse earlier than standard of care
 Risk of Disease
Risk of Disease
 Prognosis
Prognosis
 Overall survival
Overall survival
 Inform novel biomarkers
Inform novel biomarkers
 Treatment selection
Treatment selection
 Treatment response and resistance monitoring
Treatment response and resistance monitoring
 Metastatic diagnosis
Metastatic diagnosis
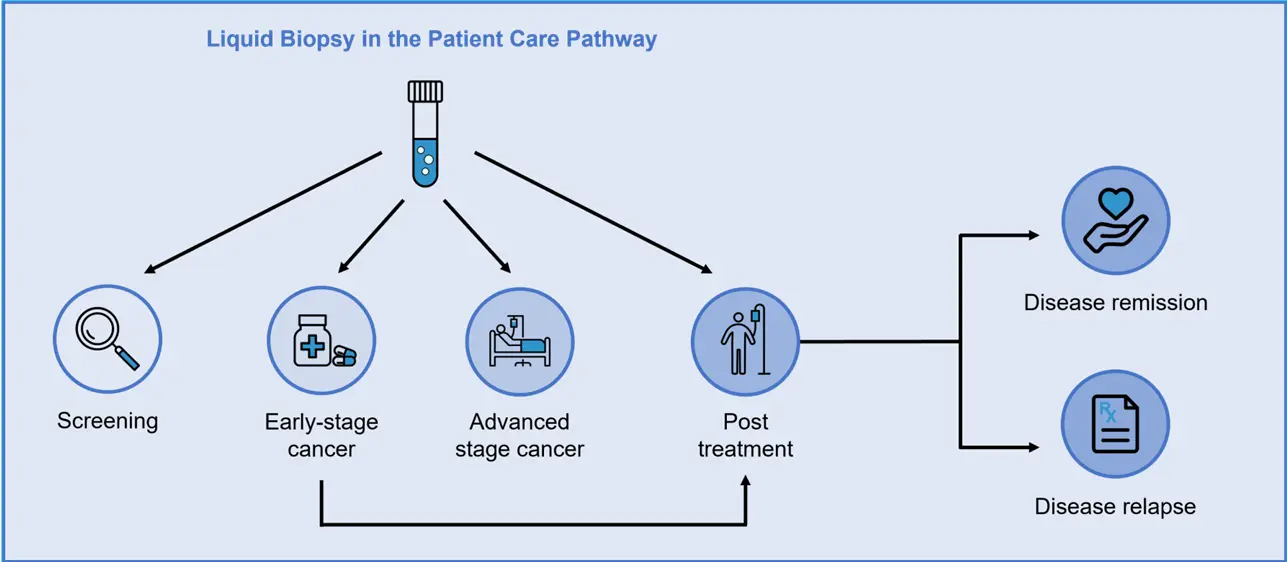
Liquid Biopsy and CTCs
Blood as liquid Biopsy:
- Minimally-invasive
- Repeatable
- Representative of cancer heterogeneity
- Versatile
- Informative
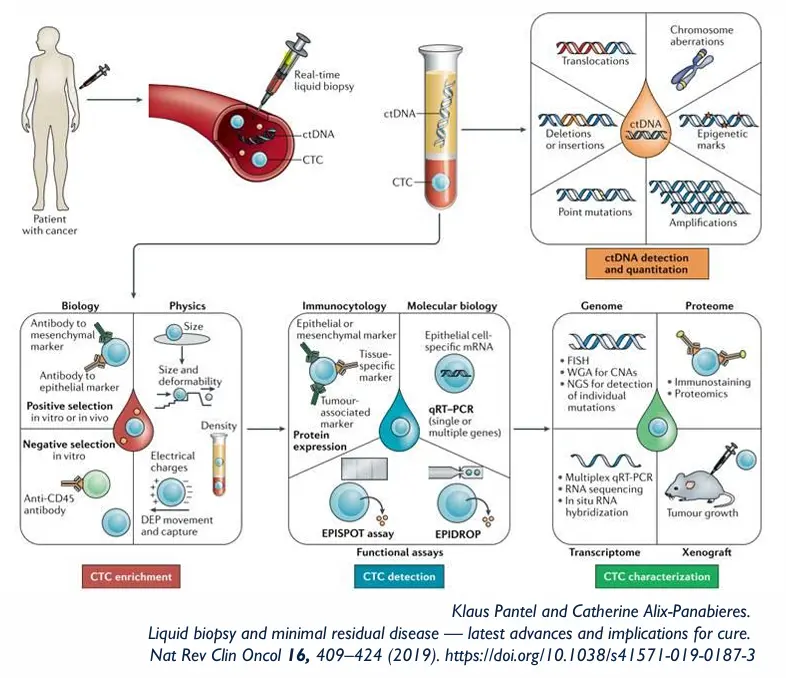

Hypothesis:
Dual analysis provides critical information not otherwise available on cancer progression and for targeted treatment selection.
“CTCs, as living cells that are active in the metastatic process, can provide prospective insight into a patient’s cancer.
In comparison, ctDNA derived mainly from dead and dying cells, provides important but historical information on patient disease." - Prof. Evi Lianidou Head of the Molecular Diagnostics Laboratory National and Kapodistrian University of Athens
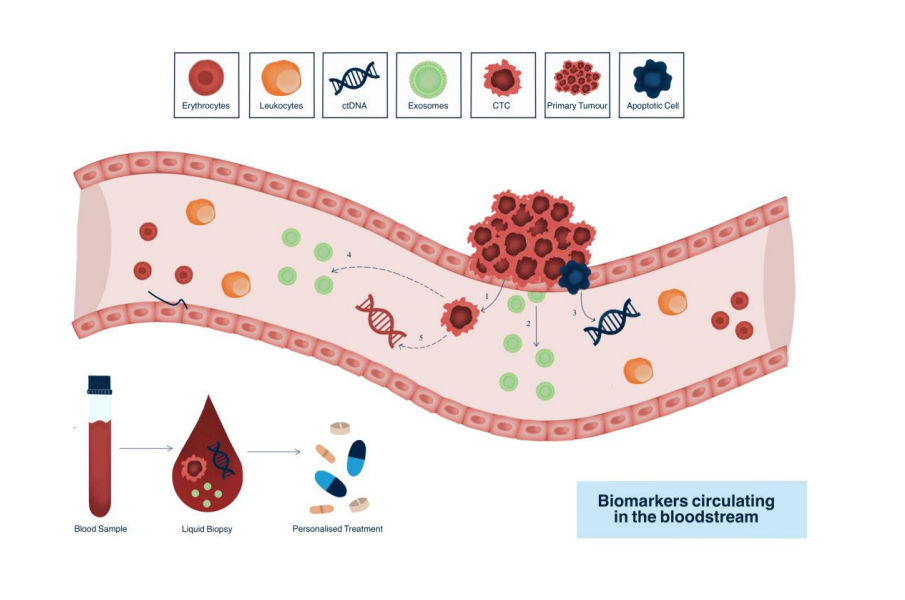
CTC and CTC clusters:
 Reflective of metastatic
tumor
Reflective of metastatic
tumor
 Source of biomarkers
(suitable for multiomics
analysis)
Source of biomarkers
(suitable for multiomics
analysis)
 Rare event: 1 CTC / 109
blood cells
Rare event: 1 CTC / 109
blood cells
Importance of upstream workflow to generate quality sample.


Intended use
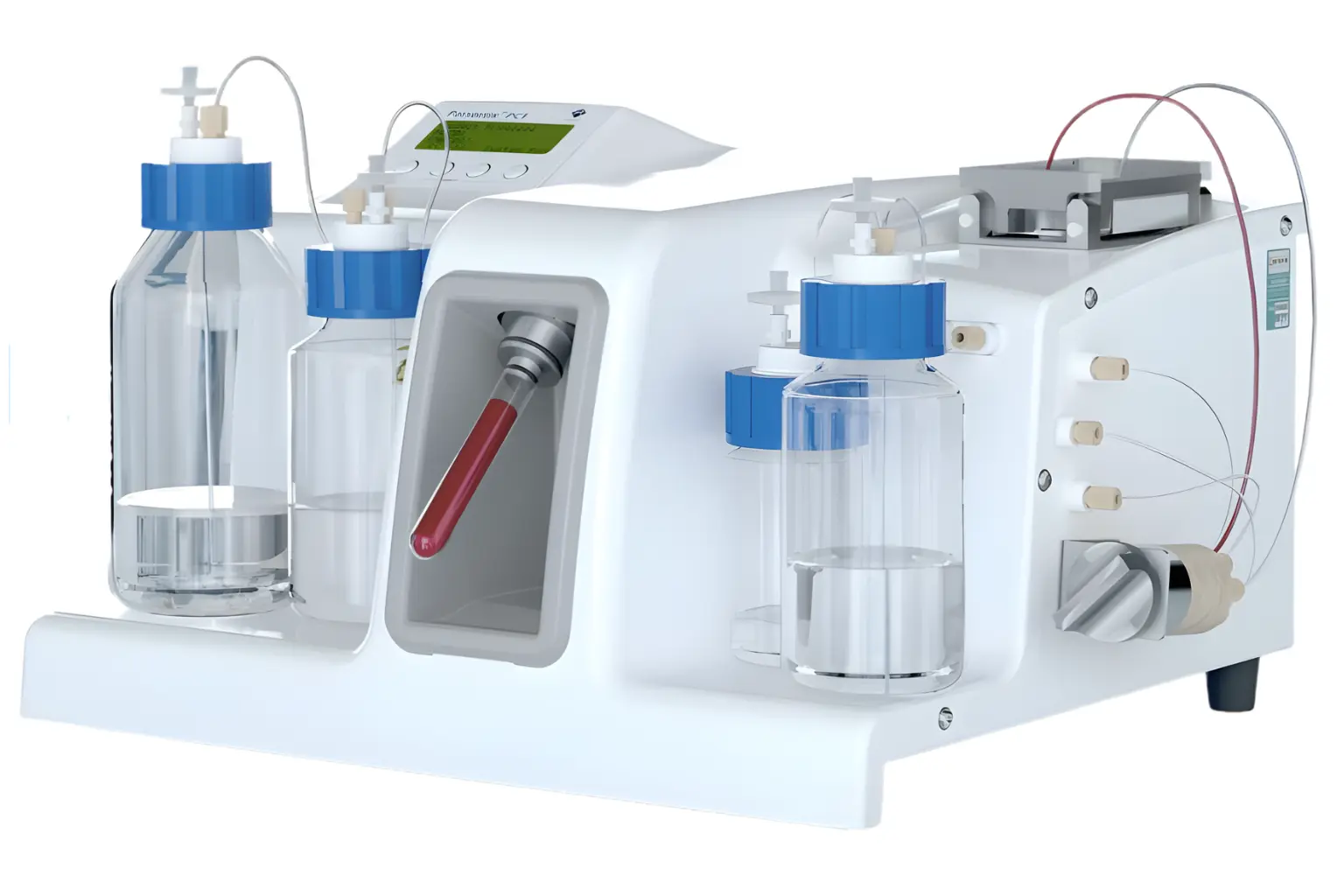
“The Parsortix® PC1 system is an in vitro diagnostic device intended to enrich circulating tumor cells (CTCs) from peripheral blood collected in K2EDTA tubes from patients diagnosed with metastatic breast cancer. The system employs a microfluidic chamber (a Parsortix cell separation cassette) to capture cells of a certain size and deformability from the population of cells present in blood. The cells retained in the cassette are harvested by the Parsortix PC1 system for use in subsequent downstream assays.
The end user is responsible for the validation of any downstream assay. The standalone device, as indicated, does not identify, enumerate or characterize CTCs and cannot be used to make any diagnostic/prognostic claims for CTCs, including monitoring indications or as an aid in any disease management and/or treatment decisions.
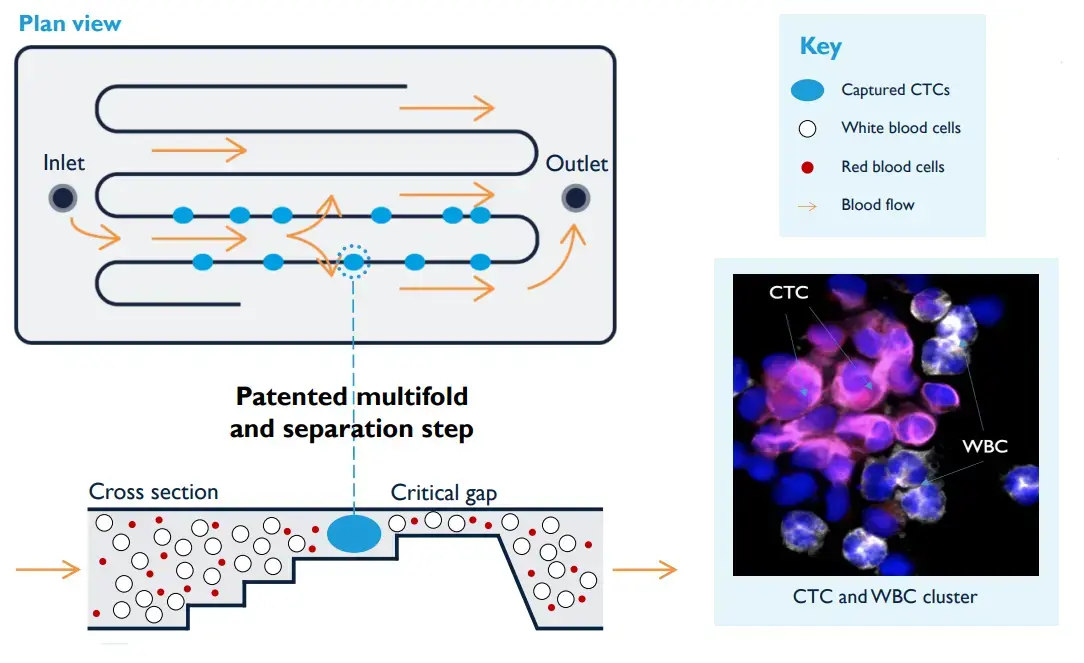
Platform technology
The Parsortix system harvests cancer cells from blood based on their larger size and lack of deformability
Other cells can be captured:
 White blood cells associated with the tumor microenvironment
White blood cells associated with the tumor microenvironment
 Megakaryocytes (frequency may relate to cancer)
Megakaryocytes (frequency may relate to cancer)
 Fetal cells from maternal blood
Fetal cells from maternal blood
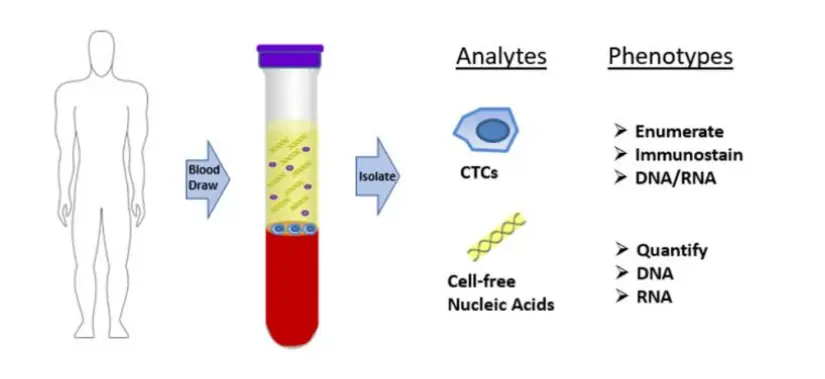
Proof Of Concept work
No CTC loss following plasma removal, prior to Parsortix enrichment.
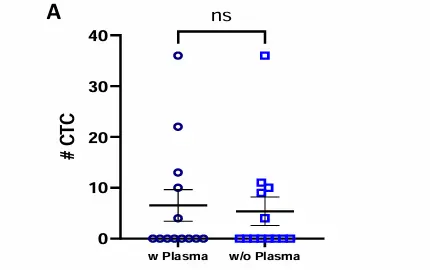
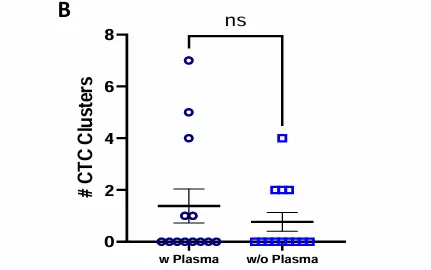
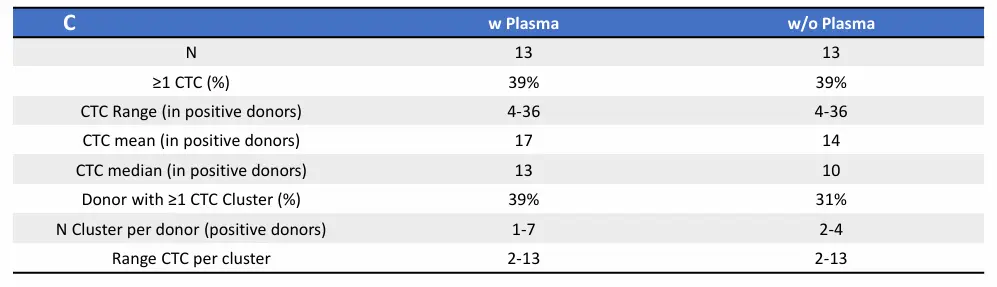
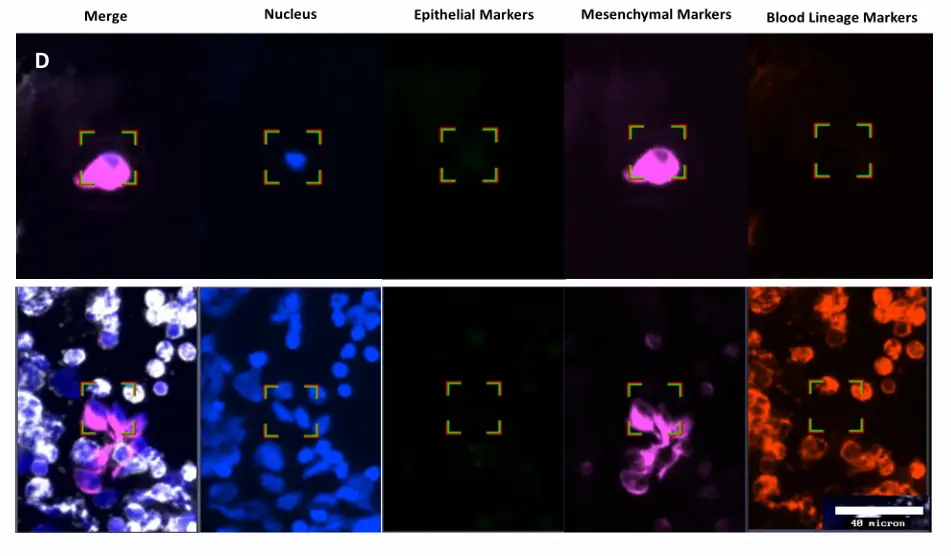
CTC detection in MBC patients blood samples processed with or without plasma. Dot plot shows mean ± SEM of the number of A) CTCs and B) CTC clusters identified in each donor per condition. No statistically significant difference was observed, ns p≥0.05, Paired T-test. C) Table showing number of donors included in each cohort (N), percentage of donors with ≥1 CTC, range, mean and median of CTCs captured within the positive group, percentage of donors with ≥1 CTC cluster, range of CTC clusters per donor, and range of number of CTCs per cluster. D) Representative images of a single mesenchymal CTC (top) and a cluster of mesenchymal CTCs (bottom). Epithelial markers (FITC) in green, Mesenchymal markers (Cy7) in magenta, Blood lineage markers (Cy5) in red or white in the merge, Nucleus (DAPI) in blue.
Can we allow for dual detection of targets/variants which may have an impact on patient care?

Can we allow for dual detection of targets/variants which may have an impact on patient care?
ctDNA and Parsortix-derived CTC-DNA can be analysed from single blood sample across multiple cancer types to detect mutations using Illumina Cell-Free DNA prep with enrichment combined with custom panel targeting Lung Cancer.
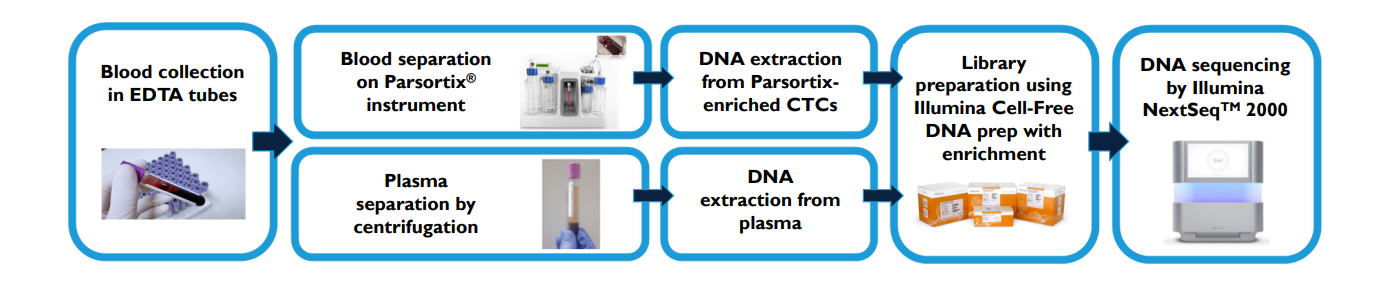
Illumina Cell-Free DNA prep with Enrichment and custom 79 gene panel enables a flexibility in the targeted genes.
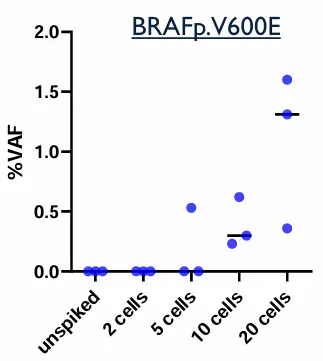
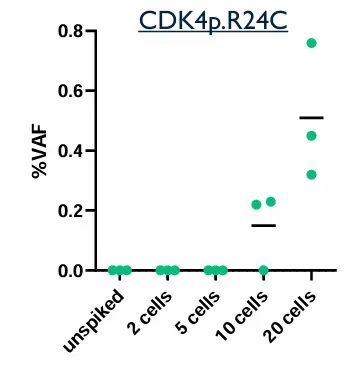
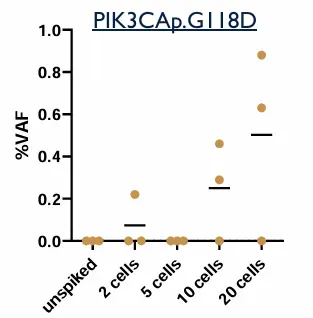
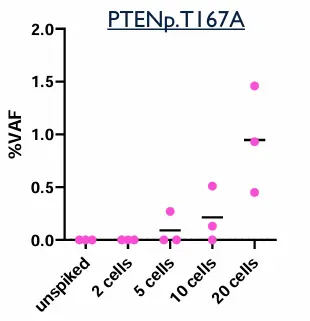
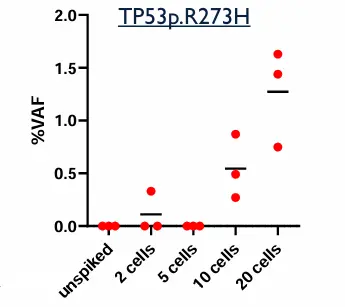
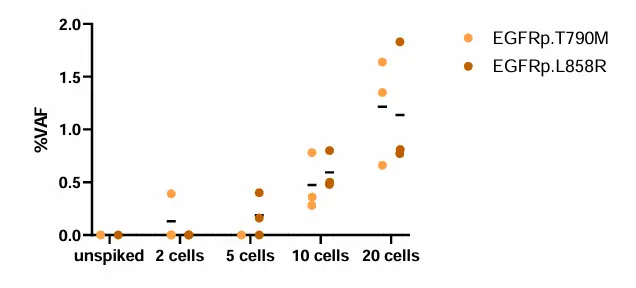
Contrived samples were used to assess the analytical performance of the assay:
 Two cell lines with known mutations, were spiked into Parsortix-enriched samples.
Two cell lines with known mutations, were spiked into Parsortix-enriched samples.
 DNA samples from CTC were extracted and processed following Illumina Cell-Free DNA prep with Enrichment procedure.
DNA samples from CTC were extracted and processed following Illumina Cell-Free DNA prep with Enrichment procedure.
Table: expected mutations to be detected in spiked samples
2 cohorts of Lung Cancer patient
- 1st cohort: 8 Lung Cancer patient samples (no treatment)
- 2nd cohort: 19 Lung Cancer patient samples (may be under treatment)

 Good quality samples
Good quality samples
 Overall, 93% of samples with at least one oncogenic gene detected
Overall, 93% of samples with at least one oncogenic gene detected
1st cohort - Lung cancer panel (n=8):
 CTC (average) = 49 and CTC (median) = 20
CTC (average) = 49 and CTC (median) = 20
 100% patients showed oncogenic mutations signal for one or more targets.
100% patients showed oncogenic mutations signal for one or more targets.
 32 genes (74 variants) seem to be specifically detected in this cohort:
32 genes (74 variants) seem to be specifically detected in this cohort:
Table: expected mutations to be detected in spiked samples
* predominant
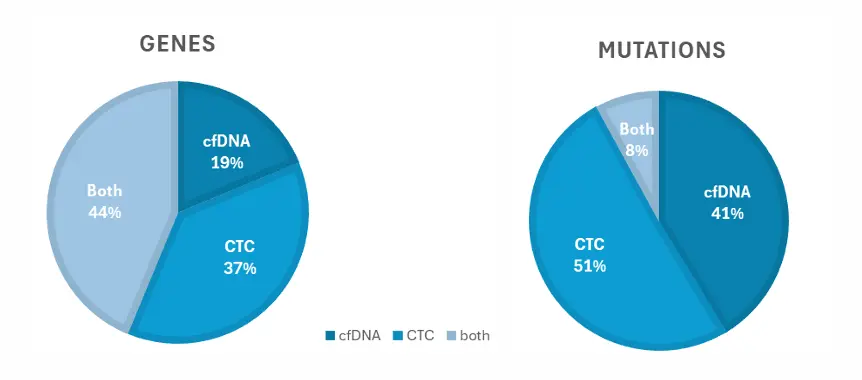
2nd cohort - Lung cancer panel (n=19):
 CTC (average) = 4 and CTC (median) = 3
CTC (average) = 4 and CTC (median) = 3
 90% of patients with at least one oncogenic mutations in CTC for one or more targets.
90% of patients with at least one oncogenic mutations in CTC for one or more targets.
 38 genes (214 variants) have been specifically detected in this cohort:
38 genes (214 variants) have been specifically detected in this cohort:
Table: expected mutations to be detected in spiked samples
* predominant
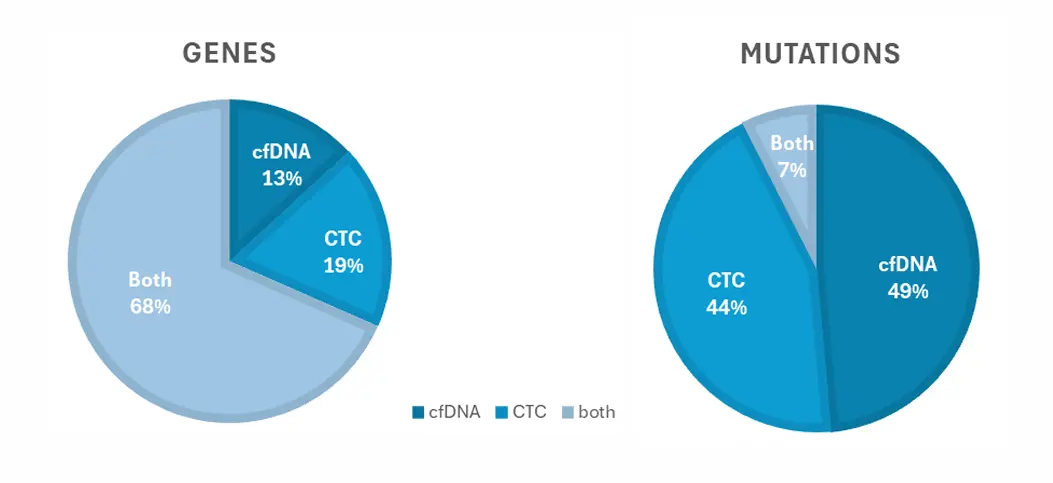
Full cohort - Lung cancer panel (n=27):
43 genes (247 variants) have been specifically detected in this cohort:
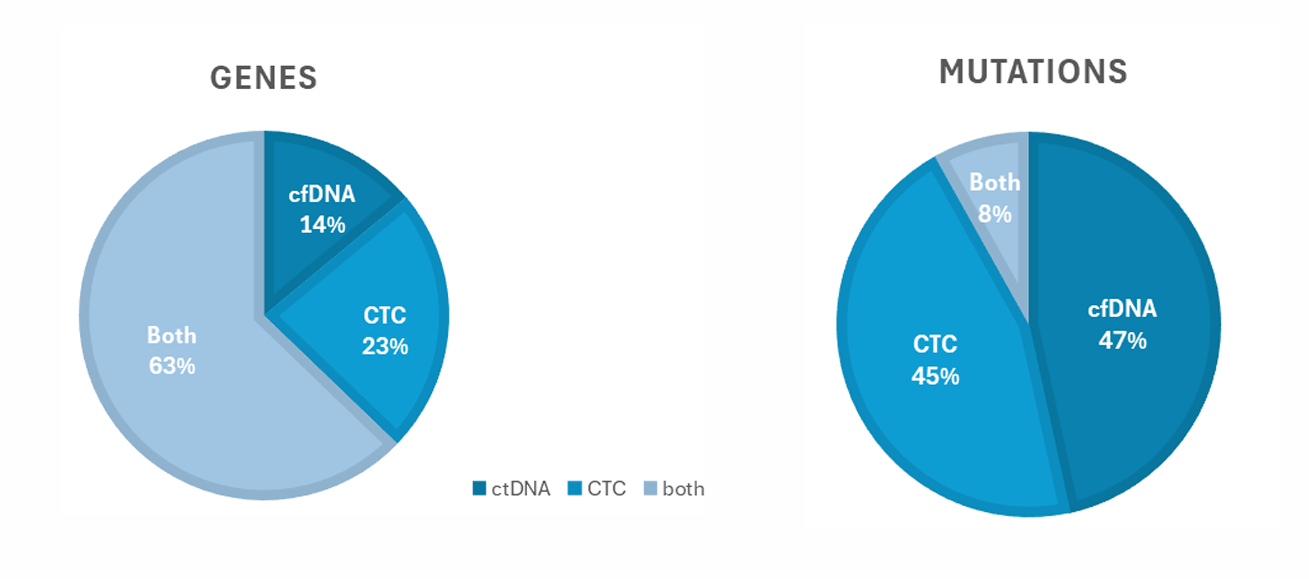
Analyzing both analytes give access to twice as many mutations, which could be investigated and/or use as “potential” biomarkers.
Validation
of patient data using another cohort of patient samples (under treatment – n=10)
-
Use of a pan-CANCER NGS panel, with blocker displacement amplification
- 20 genes (27 variants) have been specifically detected in this cohort:
Table: example of detected mutations in CTC fractions
* predominant
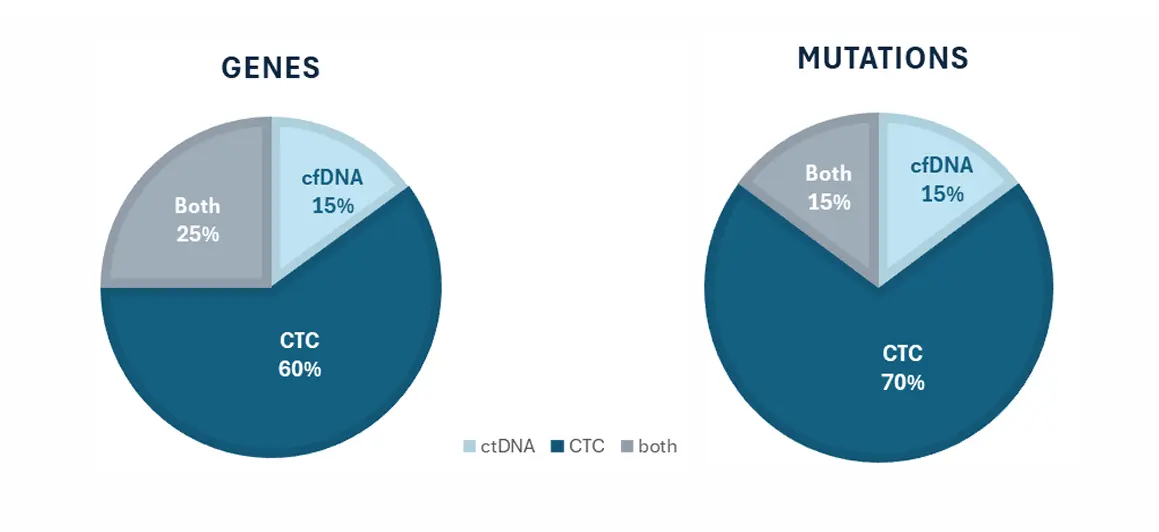
Can we allow for dual detection of targets/variants which may have an impact on patient care?

Conclusions
 Successful detection of potential actionable targets in cfDNA and CTC, from the same sample.
Successful detection of potential actionable targets in cfDNA and CTC, from the same sample.
 CTC samples provide a unique detection of actionable targets in most of the patient samples.
CTC samples provide a unique detection of actionable targets in most of the patient samples.
 This study highlights the potential for multiomic analysis.
This study highlights the potential for multiomic analysis.
 More data to come…
More data to come…
Parsortix system is allowing for CTC enrichment and DNA analysis from plasma and CTC, from thesame blood sample, with high analytical sensitivity and specificity.
Dual analysis provides the possibility to access (new) biomarkers, which may help guide patient care.
*For Research Use Only. Not for use in diagnostic procedures
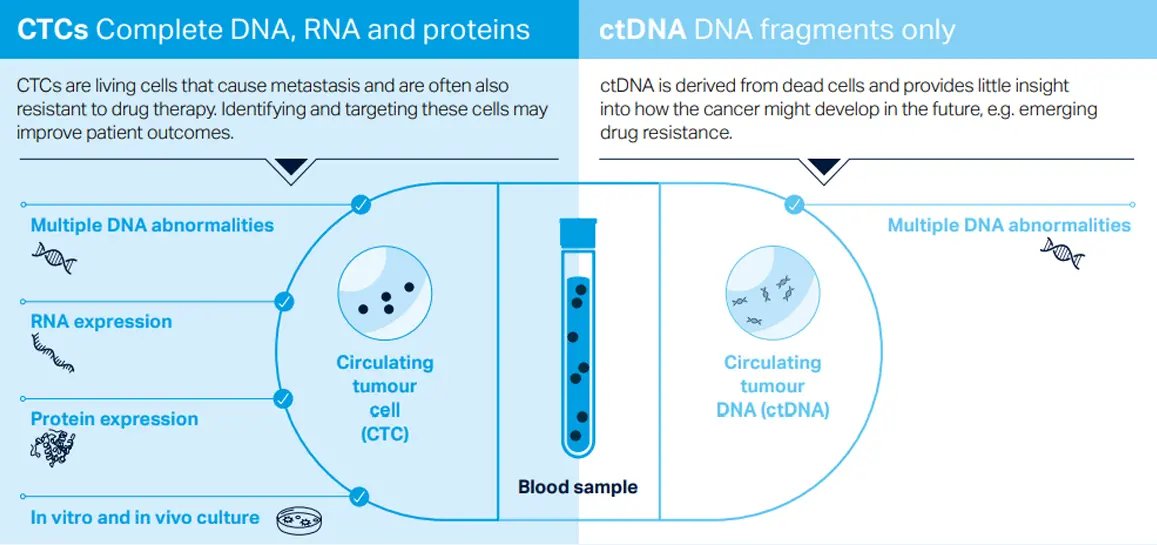
Dual analysis of CTCs and ctDNA provides complementary insights into cancer progression. CTCs reflect active and potentially metastatic disease, while ctDNA represents historical information from dead or dying cells. Together, they enable a more comprehensive understanding of tumor evolution and treatment response.
The Parsortix® PC1 system uses a microfluidic cassette to separate cells based on size and deformability. Larger and less deformable cells, such as CTCs, are retained and harvested for downstream analysis. This enables enrichment of viable cancer cells from peripheral blood.
Dual analysis can detect a wide range of oncogenic mutations, including variants in genes such as EGFR, BRAF, PIK3CA, TP53, and CDK4. In patient cohorts, up to 43 genes and 247 variants were identified. This significantly expands the number of detectable biomarkers compared to single-analyte approaches.
In the study, 93% of patient samples showed at least one oncogenic mutation using dual analysis. Additionally, combining CTC and ctDNA analysis enabled detection of twice as many mutations compared to analyzing either alone. This highlights its high sensitivity and clinical research potential.
Yes, both analytes can be extracted from a single blood sample. Plasma is separated for ctDNA analysis, while the remaining sample is processed through the Parsortix system for CTC enrichment. Importantly, no CTC loss was observed after plasma removal.
DNA from both sources is processed using Illumina Cell-Free DNA prep with enrichment and sequenced on platforms like the NextSeq™ 2000. Custom gene panels, such as a 79-gene lung cancer panel, allow targeted mutation detection. This workflow supports flexible and high-throughput genomic analysis.
While the Parsortix® PC1 system is FDA-cleared for metastatic breast cancer, the dual analysis workflow has been applied in research across multiple cancer types, including lung cancer. The methodology is adaptable to different tumor profiles using targeted sequencing panels.
CTCs are living cells involved in metastasis and can reveal ongoing tumor activity and clonal evolution. Unlike ctDNA, which reflects past disease states, CTC-derived DNA can identify emerging mutations. This makes them valuable for predicting disease progression and guiding therapy changes.
The Parsortix system enriches intact and viable CTCs, providing high-quality input for DNA extraction and sequencing. This improves analytical sensitivity and enables multiomic studies from a single sample. Platforms like CellBxHealth could further enhance such workflows by integrating advanced analysis capabilities.
Detecting a broader range of mutations increases the likelihood of identifying actionable targets for therapy. This can support personalized treatment decisions and biomarker discovery. Integrated solutions such as CellBxHealth can help translate these findings into more efficient research and clinical workflows.
Download the full Presentation in PDF format
to access detailed dual liquid biopsy workflows, including Parsortix CTC enrichment protocols, ctDNA NGS methodologies, and comprehensive mutation profiling data demonstrating enhanced biomarker detection in lung cancer samples.