Executive Summary
Choosing a CTC isolation method isn't about finding the best platform. It's about finding the one that actually serves your research. The technology you use determines not just how many cells you capture, but which cells you capture. And as you know, that distinction matters.
Antibody-based methods like CellSearch use EpCAM to grab circulating tumor cells. They're well-validated and FDA-cleared, but they systematically miss cells undergoing epithelial-to-mesenchymal transition - often the most aggressive, metastasis-driving population.
Label-free physical methods capture cells based on size and deformability instead of surface markers. They catch heterogeneous populations including EMT cells, preserve high viability for downstream work, but typically yield lower purity.
The EpCAM blind spot: In breast cancer studies, over 83% of CTC-positive patients had EpCAM-negative or mixed populations. In lung cancer, label-free methods detected CTCs in 61% of patients versus 32% for antibody-based approaches.
Why downstream applications matter: Your capture method constrains what happens next. Fixed cells can be counted and imaged. Viable cells can be cultured, tested for drug sensitivity, and used for functional studies. If your downstream application requires living cells, your platform choice isn't a preference - it's a requirement.
This guide walks through the five questions that should drive your platform decision. Answer them, and the right choice usually becomes clear.
Key Takeaways
 The cells you miss might be the ones that matter most.
The cells you miss might be the ones that matter most.
EpCAM-based capture cannot detect cells that have undergone EMT and lost epithelial markers. Head-to-head studies show label-free methods detect CTCs in 61% of lung cancer patients versus 32% for antibody-based approaches.
 Your downstream application determines your platform requirements.
Your downstream application determines your platform requirements.
Fixed cells work for enumeration. Viable cells are required for culture, drug sensitivity testing, and functional studies. This isn't a nice-to-have distinction - it's pass/fail.
 Cancer type significantly affects detection rates.
Cancer type significantly affects detection rates.
High-shedding tumors (colorectal, lung, breast) work with most platforms. Low-EpCAM cancers (renal cell carcinoma at 29% expression, triple-negative breast cancer, glioblastoma) may require label-free approaches.
 Cell loss during handling is underappreciated.
Cell loss during handling is underappreciated.
Standard laboratory techniques can lose 70% of captured cells during transfer and processing. Purpose-built consumables that retain cells through the workflow can make the difference between usable data and wasted effort.
 Purity matters most for bulk analysis; viability matters most for functional work.
Purity matters most for bulk analysis; viability matters most for functional work.
Single-cell approaches can computationally filter out contaminating cells. Bulk sequencing cannot. Match your purity requirements to your analytical method.
What questions will be answered below?
- Which cells do you actually need to capture?
- What are you going to do with the cells after you catch them?
- What cancer type are you studying?
- How much cell loss can you tolerate in your workflow?
- What does your complete workflow look like?
Why does your isolation method matter so much?
Liquid biopsy has become a practical reality for cancer research. Instead of surgical tissue biopsies - invasive, limited to a single snapshot, and sometimes impossible depending on tumor location - researchers can study cancer through a blood draw.
The premise is straightforward: tumors shed cells into the bloodstream. Catch those cells, and you can learn about the tumor without cutting into the patient.
The challenge is that circulating tumor cells are extraordinarily rare. You might have five cancer cells mixed in with billions of normal blood cells. The technology you use to find them determines not just how many you catch, but which cells you catch and what you can do with them afterward.
That's not a minor technical detail. It determines whether your research questions are answerable at all.
Question 1: Which cells do you actually need to capture?
This is the question that should come first, but often doesn't. Most researchers default to thinking about capture efficiency in general terms - "how many cells does this platform catch?" The more important question is: "does this platform catch the cells I care about?"
The EpCAM problem
The CellSearch system - the FDA-cleared gold standard for CTC enumeration - captures cells using antibodies against EpCAM, a protein expressed on epithelial cell surfaces. This works well for cells that express EpCAM. The problem is that cancer cells don't always cooperate.
When cancer cells undergo epithelial-to-mesenchymal transition (EMT), they lose epithelial characteristics and gain mesenchymal, migratory features. This transition is associated with:
- Increased invasiveness.
- Enhanced metastatic potential.
- Resistance to certain therapies.
- Downregulation of EpCAM expression.
So the cells that are most clinically concerning - the ones actively transitioning toward a metastatic phenotype - are precisely the ones that antibody-based methods struggle to detect.
How common is this problem?
The data suggests it's not a rare edge case:
Label-free capture solves this problem
Physical separation methods like the Parsortix PR1 system capture cells based on size and deformability rather than surface markers. Cancer cells are generally larger (12-25 μm versus 6-8 μm for red blood cells) and less deformable than normal blood cells. These physical properties persist even when surface markers disappear.
The Parsortix system pushes blood through a microfluidic cassette with progressively narrowing channels. The final gap is 6.5 micrometers wide. Normal blood cells squeeze through; larger, stiffer tumor cells get trapped. No antibodies determine capture. The system doesn't care what proteins are on the cell surface.
What should you ask yourself?
- Are you studying metastasis, treatment resistance, or EMT biology? If yes, you need a method that captures mesenchymal phenotypes.
- Is your cancer type known for low EpCAM expression (triple-negative breast cancer, renal cell carcinoma, melanoma)? If yes, antibody-based methods are fundamentally limited.
- Do you need to capture the full heterogeneity of circulating tumor cells? If yes, label-free approaches cast a wider net.
Question 2: What are you going to do with the cells after you catch them?
Capturing CTCs is only the beginning. What you can actually do with those cells - culture them, sequence them, test drug responses - depends heavily on how you captured them in the first place.
The fixed versus viable divide
This is the fundamental split in CTC isolation, and it's often overlooked until it's too late.
CellSearch fixes cells during processing. The CellSave preservative tubes kill the cells but preserve them for imaging and counting. You get a number. You can look at morphology. But the cells are dead.
Label-free physical methods preserve viability. By separating cells based on physical properties and using gentle pressure (the Parsortix system operates at under 0.5 psi), these systems maintain cell viability - typically above 95%.
It's the difference between count-and-describe and study-and-manipulate.
What can you do with fixed cells?
Fixed cells aren't useless - they're just limited to specific applications:
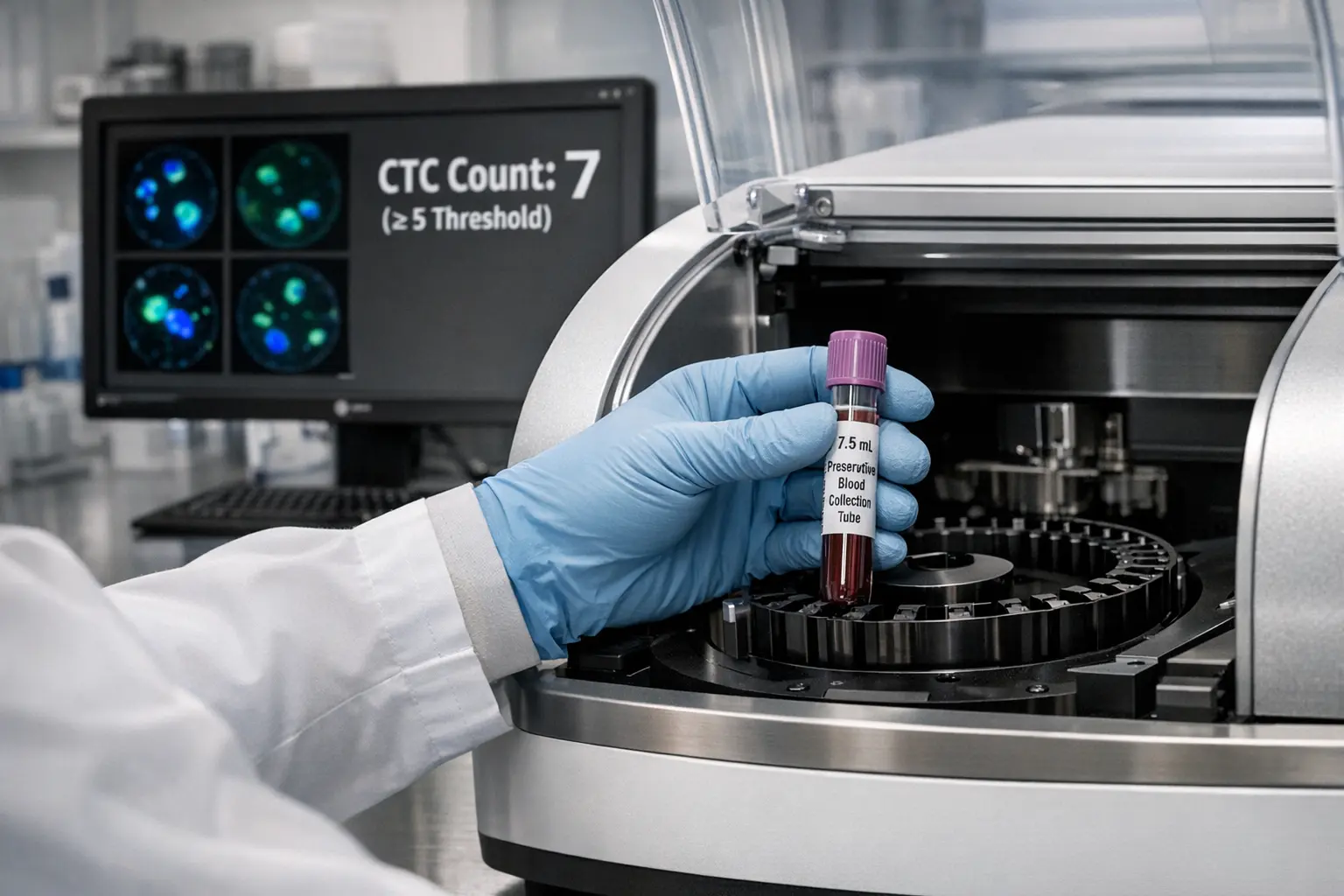
- Enumeration and prognostic assessment. The ≥5 CTC threshold in metastatic breast cancer predicts worse outcomes and is incorporated into clinical guidelines.
- Immunofluorescence imaging. Fixed cells can be stained with fluorescent antibodies to confirm identity and characterize protein expression.
- FISH analysis. Fluorescence in situ hybridization can detect gene amplifications or translocations in fixed CTCs.
- Limited DNA analysis. Possible, but yields are lower and fragmentation is higher than from fresh cells.
What requires viable cells?
If your research requires any of these downstream applications, your platform choice is already made. You need viable cells.
What should you ask yourself?
- Will you need to culture captured cells or test drug responses? If yes, you need viable cells.
- Is single-cell RNA sequencing part of your workflow? If yes, cell viability and integrity affect data quality.
- Are you primarily interested in enumeration with established prognostic thresholds? If yes, CellSearch has irreplaceable clinical validation.
Question 3: What cancer type are you studying?
Not all cancers shed CTCs equally, and not all cancers express the markers that antibody-based methods depend on. Your tumor type significantly affects which platforms will actually work for your research.
EpCAM expression varies dramatically by cancer type
The renal cell carcinoma example
Renal cell carcinoma illustrates the problem clearly. Comparative studies show the Parsortix system achieving 82-87% recovery for ccRCC cell lines versus 1-21% for EpCAM-based systems. That's not a subtle difference - it's the difference between having data and not having data.
Glioblastoma presents unique challenges
Standard ctDNA detection works in less than 10% of glioblastoma patients because the blood-brain barrier limits both CTC shedding and DNA release. This is biological, not technical - no assay improvement will fix it.
For brain tumor research, you have two practical options:
- Cerebrospinal fluid sampling - achieves higher detection rates but is more invasive.
- CTC analysis using label-free methods - CTCs detected in 77-84% of glioblastoma patients using size-based isolation.
When ctDNA isn't a viable option, CTCs become your path to liquid biopsy data.
What should you ask yourself?
- Does your cancer type have reliable EpCAM expression? If not, antibody-based methods will underperform.
- Are you working with a low-shedding tumor? If yes, maximize capture efficiency with label-free approaches and consider larger blood volumes.
- Is ctDNA detection limited for your cancer type (glioblastoma, some prostate contexts)? If yes, CTCs may be your only viable blood-based option.
Question 4: How much cell loss can you tolerate in your workflow?
You can capture CTCs successfully and then lose most of them during downstream handling. This problem is underappreciated, but it can make the difference between usable data and garbage.
Where cells get lost
Standard laboratory techniques for transferring cells to slides, concentrating samples, or moving cells between containers can result in 50-70% cell loss.
The math problem
Say you successfully capture 10 rare cancer cells from a blood sample. Standard handling techniques might lose 70% during transfer to a slide. You started with 10, now you have 3, and your statistical power is compromised.
When you're working with rare cells, the difference between 30% retention and 77% retention isn't incremental. It's the difference between having enough cells to analyze and not.
Purpose-built consumables solve this problem
The CellKeep Slide system addresses cell loss by concentrating captured cells into a small, defined area rather than spreading them across a large surface. The system uses a proprietary wicking cap that removes liquid without disturbing deposited cells.
The result: 77% cell retention versus 30% for standard methods.
This also reduces the volume of expensive staining antibodies needed - you're concentrating cells into a smaller area, so you use less reagent.
What should you ask yourself?
- How many CTCs do you typically capture per sample? If the number is low (and it usually is), every cell matters.
- What's your current cell retention rate through the full workflow? If you don't know, you should find out.
- Are you using standard lab consumables for rare cell handling? If yes, you're probably losing more cells than you realize.
Question 5:
What does your complete workflow look like?
CTC isolation doesn't happen in isolation. Your platform needs to integrate with sample collection logistics, processing timelines, downstream analysis, and practical constraints like staffing and throughput.
Sample stability and logistics
CellSearch's 96-hour stability window is a genuine operational advantage for multi-site studies with centralized processing. Blood can be drawn at a clinic, shipped overnight, and processed at a centralized lab.
Label-free methods require more immediate processing unless you use preservative tubes. For labs processing their own samples, this is less of an issue. For multi-site studies, it requires more careful logistics.
Purity considerations
Size-based filtration isn't perfectly specific. Some white blood cells (especially monocytes, which can reach 20 μm) may contaminate the output. CellSearch's antibody-based selection produces cleaner populations.
Whether this matters depends on your downstream application:
- For enumeration: Largely irrelevant - you're staining and counting anyway.
- For bulk sequencing: Contaminating cells contribute their own signal and can skew results.
- For single-cell analysis: You can computationally identify and exclude contaminating cells after the fact.
Characterizing what you caught
After capture and concentration, you need to identify and characterize your CTCs. The Portrait+ immunofluorescence kit provides four-channel staining that distinguishes:
This combination detects cells across the EMT spectrum - not just whether cells are present, but where they fall on the epithelial-to-mesenchymal continuum. That's information you can't get from systems that only capture (and therefore only characterize) epithelial phenotypes.
What should you ask yourself?
- Is this a single-site study or multi-site with centralized processing? Sample logistics matter.
- What's your processing capacity? Can you handle same-day processing, or do you need extended stability?
- Do you need to characterize EMT status? If yes, your staining panel needs to include mesenchymal markers.
- What level of purity does your downstream analysis require?
How do these questions point you toward a platform?
The right CTC isolation method depends on your answers to these five questions. Here's how different answers point toward different platforms:

Choose antibody-based methods (CellSearch) when:
- You need validated prognostic enumeration with established clinical thresholds.
- Your study requires multi-site sample collection with centralized processing.
- You're specifically interested in EpCAM-high epithelial CTCs.
- Maximum standardization and automation are priorities.
- Insurance reimbursement is a factor for clinical applications.
Choose label-free physical methods (Parsortix) when:
- You need to capture heterogeneous CTC populations, including EMT cells.
- Downstream applications require viable cells.
- You're studying cancers with low EpCAM expression.
- You want to detect the broadest possible CTC population.
- You need flexibility in sample volume.
- EMT biology or mesenchymal phenotypes matter to your research.

Consider the complete CellBx Health workflow when:
- You need label-free capture that preserves viability.
- Cell retention through the workflow is critical.
- You want to characterize both epithelial and mesenchymal phenotypes.
- You're working with rare cells where every captured cell matters.
Your platform choice is a research design decision
The platform you choose for CTC isolation isn't a neutral technical decision. It determines which cells you can catch, what you can do with them, and ultimately whether your research questions are answerable.
Start with what you need to learn. Identify which downstream applications will answer your questions. Then choose a capture method that supports those applications - not the other way around.
For researchers whose questions require capturing the full heterogeneity of circulating tumor cells, including the aggressive EMT populations that antibody methods miss, label-free approaches aren't just preferable. They're necessary.
And for researchers working with rare cells where every captured cell matters, the complete workflow - from capture through concentration through characterization - determines whether your CTC research succeeds or fails.
Yamato Life Sciences offers the CellBx Health liquid biopsy system, including the Parsortix PR1 for label-free CTC capture, CellKeep Slide for high-retention cell concentration, and Portrait+ immunofluorescence kit for CTC identification and EMT characterization.